Enhancing the power of super-resolution microscopy
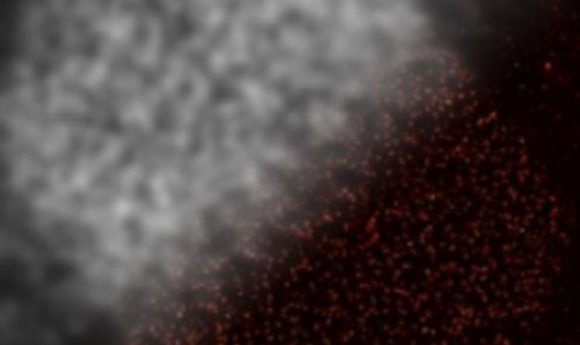

Novel DNA aptamer fluorescent markers could enable the visualization of entire protein networks in high resolution.
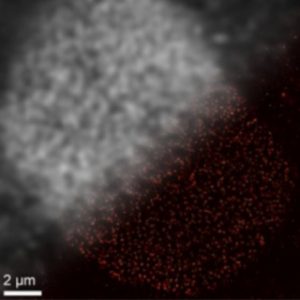
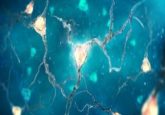
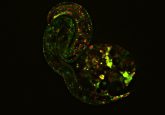
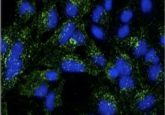
Looking from top-left to bottom-right, you can see the difference with traditional imaging (top-left) in comparison to imaging with DNA-PAINT (bottom-right). Credit: Ralf Jungmann, Ludwig-Maximillians-Universitaet.
Researchers from the Ludwig-Maximillians-Universitaet (Munich, Germany) have utilized DNA aptamers that are covalently attached to fluorescent markers in order to locate target proteins with complementary DNA tags. This technique makes it possible to image how molecules are distributed in a single cell at super resolution.
Previous fluorescent markers for target proteins have been too large and this has been a key factor slowing down the progression of super-resolution microscopy. Therefore, the team decided to downsize the markers by modifying DNA aptamers – single-stranded DNA molecules that can fold into 3D shapes and specifically target different proteins.
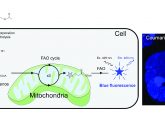
They combined the DNA aptamers with fluorescent labels, developed protocols for labeling of fixed and live cells, and termed the technique DNA-PAINT. The study, published in Nature Methods, has also shown that the resolution attained with these new markers alongside super-resolution microscopy is an improvement on current super-resolution methods.
“We expect that the new method will provide a significant boost for super-resolution microscopy, particularly with respect to its range of application in biology,” commented Ralf Jungmann, a professor of experimental physics at the Ludwig-Maximillians-Universitaet and co-author of the study.
Jungmann’s goal is to enable the visualization of whole protein network interactions at a single-cell level.
“We will be able to address biological and biomedical questions that have thus far been experimentally inaccessible.”