Decoding the secrets of totipotent cells
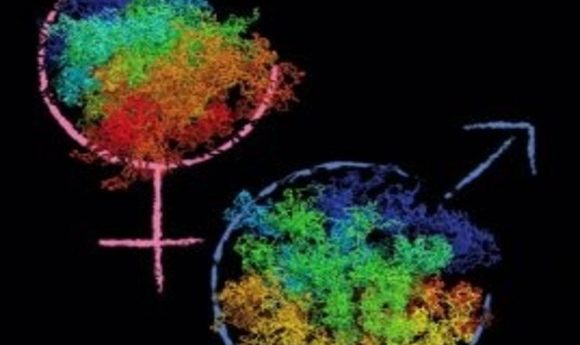
A new method reveals critical differences in the packaging of maternal and paternal genomes in zygotes.
Image courtesy of Kikue Tachibana-Konwalski
How a totipotent cell retains the potential to become any of the 200+ cell types in the human body is a longstanding mystery in biology. Researchers believe that these cells must have a unique pattern of global gene expression that accounts for this remarkable property. But because of technical limitations, it has been challenging to understand key aspects of gene regulation, such as how enhancers find promoters when they are separated by hundreds of kilobases of sequence.
Now, a new method allows researchers to look at the 3-D organization of the genome in single cells and pin down which DNA sequences are next to each other inside zygotes. The study, appearing in Nature, reveals that the maternal and paternal pro-nuclei in zygotes are packaged differently, leading to differences in gene activity that may reveal the biological basis of totipotency.
Fifteen years ago, researchers developed the “chromosome conformation capture” (3C) method to study how the genome is organized in cell populations. Briefly, 3C involves fixing the cells with formaldehyde, digesting their chromatin with a restriction enzyme, and then ligating the DNA. During the ligation step, a DNA fragment can reconnect to its original strand, or it can connect to a piece of DNA that was far away in terms of sequence but spatially close. Typically, researchers add a biotin “adaptor” to the DNA ends before ligation, which enriches for informative fragments in the ligation step. By analyzing these ligation junctions, researchers can pinpoint which sequences are found next to each other on average in a population of cells.
Kikue Tachibana-Konwalski of the Institute of Molecular Biotechnology in Vienna and her student Ilya Flyamer (now at the University of Edinburgh) developed an improved method that produced tens to hundreds more contacts per cell. “The key change in our method was not adding the biotin after chromatin digestion. Since the DNA fragments have sticky ends during ligation, it works more efficiently, and we can capture many more interactions from a single nucleus,” explained Flyamer.
In a zygote, maternal and paternal pro-nuclei are separate, so Tachibana-Konwalski and another student, Johanna Gassler, extracted the pro-nuclei and analyzed them separately. “We found that the global organization of zygotic maternal and paternal genomes is fundamentally different from other cells and discovered unexpected differences in chromatin structure,” Tachibana-Konwalski said.
The paternal genome is organized into compartments, meaning that regions of transcriptional activity and regions of transcriptional repression are spatially segregated in the nucleus. The maternal chromatin, however, seems to have no such segregation. These differences are functionally important; the paternal genome starts transcribing during the first cell cycle, but the maternal genome doesn’t begin until the two-cell stage. “So far, every cell type analyzed has always had compartments. The maternal genome is the first that does not, and we think that this may be related to the state of chromatin reprogramming,” said Tachibana-Konwalski.