Streamlining image analysis for mitotic cells with AI
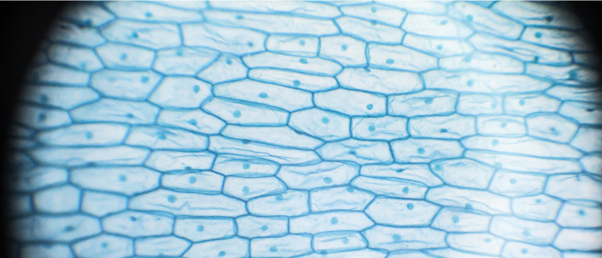
Researchers developed a deep-learning model for artificial intelligence (AI) to recognize mitotic cells, which is easy-to-use, easy-to-train, and aimed at non-data scientists.
Advancements in microscopy over the centuries have provided scientists with higher quality images and a more detailed understanding of biological processes, such as cell division. Scientists can follow mitosis by studying microscopy images of chromosomes; however, images continue to be analyzed manually, which is a time-consuming process.
With the introduction of automation to microscopy in recent years, more images can be taken in a shorter period meaning scientists can track the pathway of mitosis in more detail. However, with more images comes more analysis, which can easily be more than 1,000 images for large specimens using automatic acquisition methods. Analyzing such vast quantities of images is especially tedious when looking at mitosis in plants with a large variety of chromosome structures. Clearly, automating this process would be hugely beneficial to researchers.
Now, researchers from Okayama University (Japan) have developed a deep-learning AI model to classify images for several plant species. This is not the first of its kind; however, it is the first that is easy-to-use for non-data scientists. Kiyotaka Nagaki, who led the research, explains that “classifying images using AI usually requires a high level of computer knowledge. What we did is build AI models on a McIntosh computer with the Create ML app suitable for our own image samples. Moreover, the AI can be trained to become an order-made image classifier for any variety of images that suits one’s purpose.”
 Could artificial intelligence predict your risk of stroke?
Could artificial intelligence predict your risk of stroke?
Research suggests that AI could be used to predict atrial fibrillation-related stroke risk in patients.
The group used a variety of images containing mitotic cells from plants to train the deep-learning model to recognize where cells are undergoing mitosis. Nagaki describes this AI model as “a deep learning sorter that anyone can use” because it has been designed for those who don’t have a computational background.
Following the learning period, the model was tested using images from different plant species, which were successfully classified. Additionally, it was able to identify cell division in tissue sections meaning this technology could be applied to other biological images.
“There are more trivial classifications in our lives than one might imagine. Automating such classifications by entrusting them to an AI can not only eliminate fluctuations caused by individual differences but also save many valuable research hours,” says Nagaki. “Streamlining such trivial classifications make extensive image-based studies more reproducible and reliable.”
Computer models that allow researchers to analyze and interpret their data, without having to become experts in computational biology or bioinformatics, present a huge time-saving opportunity. By allowing researchers to focus on their time and intellect on the question at hand, models like this can greatly accelerate the pace of a projects process and scientific discovery.