The CRISPR answer to antibiotic resistance
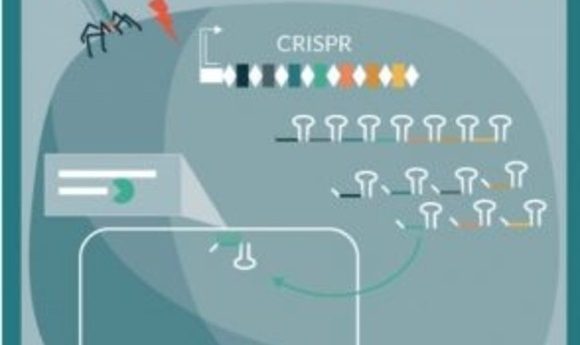
Long before CRISPR became the genome-editing superstar it is today, it played an important role in protecting bacteria from invading phages. Now researchers are bringing it—along with its new skillset—back to its bacterial roots to solve the rising problem of antibiotic resistance and offer a realistic means for microbiome farming.
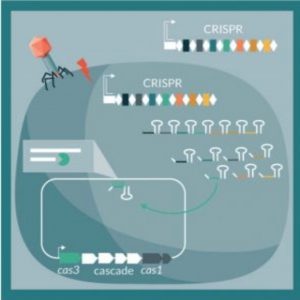
Researchers are redirecting bacterial CRISPR systems to attack their own host chromosomes (3).
Over the last five years, the CRISPR-Cas system has risen to fame as researchers developed it into a powerful and precise genome-editing tool. Since its introduction by Jennifer Doudna and Emmanuelle Charpentier in 2012, CRISPR gene editing has restored hope for gene therapy, expanded our ability to create animal models of human disease, showed promise for treating viral infection, and simplified gene editing tasks so much that nearly everyone can perform these experiments. While there have been bumps in the road, CRISPR so far is living up to its hype.
It wasn’t always clear, however, that CRISPR would become such a superstar. In fact, it hails from a rather humble beginning; its native role is protecting bacteria from invading phages. “I remember first learning about CRISPR seven years ago when little was known about its basic mechanisms, let alone its biotechnological potential,” said Chase Beisel from North Carolina State University. “I was struck by how the system managed to differentiate between self and non-self and how simple it would be to trick the system into targeting the bacterial genome as if it was foreign genetic material.”
Now, researchers who knew CRISPR before it was famous are subverting its original function by using it to kill bacteria instead of protect them. The approach looks like a promising tool in the fight against antibiotic resistance and offers more precision than probiotics for altering the human microbiome to improve health.
Returning Home
Rodolphe Barrangou, also from North Carolina State University, has studied CRISPR for over a decade. His work was instrumental in revealing its function in bacteria and archaea, so when the system was repurposed as a genome editing tool, he was eager to use it. He and his team tested CRISPR for cutting a variety of sequences, but when they targeted the bacterial chromosome, most of their cells died. “Death is the primary outcome; editing is a side-effect—not vice versa—since DNA repair is not as advanced in bacteria as in eukaryotes,” he said. “We then realized we could use self-targeting to kill, rather than edit, bacteria.”
When a bacteriophage infects archaea or bacteria, the CRISPR system stores pieces of phage DNA as a type of immune memory. The cell can then transcribe guide RNA from the stored sequence, pair it up with a Cas enzyme, and set the duo loose to surveil the cell for matching invaders. When the RNA finds its match, Cas cuts the phage DNA, preventing it from replicating.
Barrangou’s team reasoned that if they could deliver specific guide RNAs to bacteria, they could provoke the native CRISPR system to cut plasmids carrying antibiotic resistance genes to prevent their spread or even to cut up the cell’s own chromosome, selectively eliminating certain bacteria from a population. In 2014, he, Beisel, and their colleagues provided a proof of this principle in the journal mBio, where they showed that they could induce cell death either with native CRISPR systems or by importing the CRISPR components into cells, and that cell death resulted regardless of where in the genome they targeted. Their approach enabled them to specifically eliminate certain bacteria from mixed cultures and titrate the level of cell death (1). With these results in hand, the team saw potential for developing this CRISPR-based method as a new class of antibiotics.
“The beauty of CRISPR so far is that it can attack and kill antibiotic-resistant bacteria as easily as antibiotic-susceptible bacteria. The other benefit is that CRISPR can be targeted to only the pathogens, potentially preserving our native bacteria and their health benefits,” Beisel explained.
Resisting Resistance
To deploy this CRISPR system as an antibiotic, researchers have introduced a few different delivery options, with bacteriophages showing the most promise. “The challenge with bacteriophages though is that they are highly complex machines that have not been extensively engineered. In many regards, we’re still learning how to modify bacteriophages so they reliably package our DNA of interest and deliver it to a particular set of bacteria,” Beisel said. His team is now working on modifying bacteriophage tail fibers, which recognize and bind target bacteria, to expand the range of potential bacterial targets for CRISPR antibiotics.
Many lytic bacteriophages bring with them innate microbicidal properties, a characteristic Dave Ousterout from Duke University plans to incorporate into his designs for CRISPR-based antibiotics to boost therapeutic efficacy. While lytic phages can kill cells on their own and have excellent safety profiles, they need to replicate within a host cell to exert their effect and they have a limited host range that isn’t easy to alter. Loading them with CRISPR circumvents the need for replication to kill the cell and avoids regulatory concerns that come from using a propagating—and potentially mutating—phage.
“With CRISPR-weaponized phages, these therapies can be developed and optimized against more traditional pharmacokinetic parameters, such as persistence, time, and concentration at the site of infection,” Ousterout explained.
While CRISPR carried in phages seems ideal for treating bacterial infections, all antimicrobial agents exert strong selective pressures, raising the possibility that bacteria will develop resistance. Since CRISPR is so versatile, however, researchers may be able to keep ahead of the changes and quickly redesign new generations of CRISPR antibiotics to outwit bacterial evolution.
“We are highly confident that CRISPR-Cas3, owing to its ability to target many sites in the genome at the same time, can avoid potential resistance mechanisms,” Ousterout said. For killing cells, Cas3 is a better option than Cas9, which is commonly used in CRISPR genome editing tasks; Cas9 creates double strand breaks that cells can repair, whereas Cas3 cleavage is irreversible. “Our approach to developing suitable bacteriophage vectors relies on using a cocktail of phages that target bacteria in different ways. Together, these approaches will mitigate the chance of rapid resistance.”
Microbial Farming
While developing a new class of antibiotics is a high priority for human health, this technology may extend far beyond treating bacterial infections. In recent years, the microbiome has become almost as notable as CRISPR itself, and has proven to affect human health in surprising ways, from metabolism, cardiac function, and pain to neurodegeneration and mental health.
Many researchers see the potential of altering the microbiome to improve health and are keen to develop ways to do so. Fecal transplants and probiotics have proven helpful in limited cases, but these treatments can be temporary since selective pressures that built certain microbiomes in the first place are likely to push the bacterial population back to its original balance. “This technology has the potential to enable highly-selective removal of select species in complex, polymicrobial environments, such as the microbiome,” Ousterout said.
This level of precision facilitates much deeper studies of microbial communities since individual members can be selectively eliminated to determine their role in the overall bacterial population. It also offers a realistic mechanism for microbial farming—selectively cultivating healthy microbiomes.
Barrangou, Beisel, Ousterout, and several other colleagues recently co-founded Locus BioSciences to develop narrow spectrum antimicrobials for addressing outbreaks of antibiotic resistant infections. The company is currently developing a phage-delivered CRISPR system for eliminating C. dfficile without harming other gut microbes. “We think that this will allow the normal microbiome to be restored and provide protection against recurrent C. difficile infections,” Ousterout said. “With this proof-of-efficacy in hand for a novel narrow spectrum antimicrobial, Locus would move to apply this technology to other bacteria in the microbiome that may cause a variety of human diseases, such as colitis, diabetes, and potentially even neurological disorders.”
These goals may come earlier than expected. In June, Barrangou presented at the CRISPR 2017 conference in Big Sky, Montana that the new CRISPR antimicrobial works and is tolerated well in lethal infection and bioburden animal models.