Guohao Dai: 3D bioprinted glioblastoma models and 3D imaging
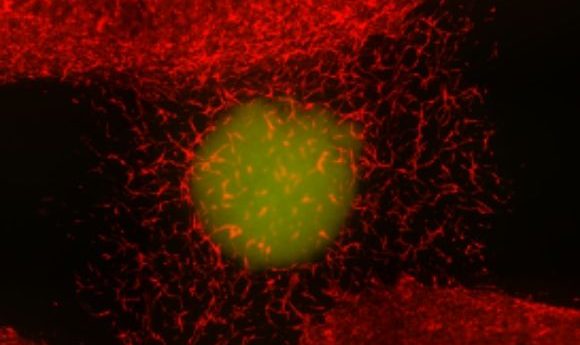
We speak to Guohao Dai about his recent paper, which covers 3D printing techniques for creating more accurate glioblastoma tumor models and improvements to the imaging techniques used to study them.

BioTechniques Digital Editor, Tristan Free, catches up with Guohao Dai, Associate Professor of Bioengineering at Northeastern University (MA, USA), to discuss his recent work making improvements in glioblastoma modeling and imaging, using 3D printing techniques.
Can you tell us about the technique that you recently developed for 3D printing tumor cell models?
We recently published this in a paper for Science Advances. We developed a 3D bioprinted glioblastoma tumor model, which is the deadliest form of brain tumor. The 3D bioprinting model was developed to mimic the extracellular microenvironment in which we could grow the glioblastoma tumor cells. We also developed an imaging technology capable of imaging these tumor models that are thicker than typical models, allowing us to image several millimeters deep. This combination of novel techniques – using fluorescence imaging to study the bioprinted tumor models – meant that we were able to reconstruct the 3D shape of the tumor and then see how it grows in real time.
How did 3D printing help with the process of developing these 3D models as opposed to other commonly used techniques to produce 3D tumor spheroids or organoids?
As you say, you can use a variety of techniques to produce 3D tumor models. In our case, we used 3D bioprinting because we can print a 3D vascular channel together with the 3D tumor model, so the tumor model is actually being perfused with vascular channels. It is much easier to form these vascular channels accurately with 3D bioprinting.
What does this method allow you to study that you wouldn’t have been able to in animal models, 2D cell cultures or organoid models produced using different techniques?
First of all, the animal model is currently the gold standard for conducting brain tumor research. You inject the patient’s glioblastoma tumor cell into the mouse brain – this is called intracranial transplantation – then you can study how the tumor invades the brain and monitor its response to drugs. This is, however, a very expensive and lengthy process. During the 6 months after you inject the tumor cells into the mouse brain, you cannot tell much about what’s happening in the brain and the tumor development until you open the cranial window. Once the cranial window is open you can observe what is happening there, but there is no real-time monitoring of how the tumor interacts with the brain tissue or how they grow.
By doing this 3D tissue modeling in vitro, we will be able to visualize the tumor cell growth in real time. We can monitor them daily to see how they respond to treatment. The other thing that’s involved with our 3D modeling is that we have incorporated vasculature channels to allow for vascular perfusion. We can allow the channel of the tumor model to be perfused with a drug for over 2 months, which is a very long time. This means that you can test long-term, which is more reflective of the treatment in real life. This is difficult to do with other models, therefore, that’s the key benefit of this 3D model.

Evolution of 3D cell culture cancer models
Check out this webinar, presented by Keith L. Ligon of Brigham and Women’s Hospital (MA, USA) and David Tuveson of Cold Spring Harbor Laboratory (NY, USA), discussing the evolution of 3D cell culture cancer models and documenting their recent advances and their potential impacts in the future of cancer research.
Could you tell us more about the imaging technique that you developed? How did it allow you to image deeper into these models?
When you develop a 3D model, usually it is very thick, at least a few millimeters deep. That presents very significant challenges to 3D imaging. For example, if you are using a confocal microscope, you only image to less than a millimeter, a few 100 microns deep. Confocal microscopes are also not ideal for constructing 3D images as it takes a few hours to reconstruct. Due to the long time, the sample is photobleached, which causes some photo damage to the cells and to the model. That’s a lot of interruption if you do that every day with a few hours of imaging time and it will also disrupt your image.
The imaging technique we developed uses a laser to quickly scan the tumor, capturing and tracking all the scattered light from the tissue and then reconstructing the image using a mathematical model that we developed, which can identify the original location of the scattered photons. That is all carried out very quickly, we scan for just a few minutes then we are done and we can take the sample back to the incubator. This means there is very little damage or disruption to the cell itself. Of course, the gain in speed and depth of imaging does come at the cost of losing resolution. We cannot image to the cellular level but we can image to few hundred microns resolution, so we are able to visualize that the tumor has overall growth.
What were some of the most exciting findings that you discovered in this study?
Once we developed a 3D model and 3D imaging, it allowed us to do some further research, however, in this paper we just did an initial study to see how this tumor responds to very typical chemotherapy, temozolomide, which is the most commonly-used chemotherapy for glioblastoma.
We tested the temozolomide in the tumor model and found that over 2 months of treatment, the tumor initially shrank. However, after the initial remission they then grew more aggressive populations of cells that were resistant to the temozolomide. Over this time, the chemotherapy did not kill the tumor, which indicates a very strong drug resistance of the glioblastoma to the chemotherapy.
Next, we want to use these models to study tumor cell dormancy; how the dormant population of the glioblastoma can resist radiation and chemotherapy.
What were some of the complications or challenges you faced when you developed these techniques and how did you overcome them?
I think the tumor model itself is still a significant challenge so far. Right now, we are only able to include some of the cells in the 3D tumor model that would be present in an in vivo tumor. In this case, we have a brain tumor cell, but we also have a brain vascular endothelial cell. We have included a parasite and the brain astrocyte, but there’s a lot of other complicated structures like neurons and oligodendrocytes and microglia in the brain.
The other thing to consider is the extracellular matrix in the brain environment. We are using hydrogels, but it is still far from an actual brain tissue in terms of the chemical composition and the biomechanical property. Therefore, I think the tumor model is still far away from actual real human representation.
Another challenge is mimicking the blood–brain barrier (BBB). At this moment, the drug will go through the bioprinted vascular channel and then cross the endothelial barrier which, in this case, is the brain endothelial cell. The challenge involving the brain endothelial cell in our bioprinted channel is they are not as tight as the BBB in humans, so there is still a significant challenge in how to recreate the extremely tight barrier.
Where do you see this project in the next 5 years? What impact do you think it can have on the field of glioblastoma study?
The dormancy study that we’re working on is to identify the dormant cell population in the glioblastoma tumors and identify the underlying molecular mechanisms of how they confer drug resistance. With glioblastomas, typically you treat the initial tumor and they often come back, indicating a very strong dormant population. I think in the next phase of the research we should be able to uncover some of the mechanisms of the dormancy and then be able to develop drugs that specially target the dormancy of the glioblastoma, hopefully giving patients much longer survival.
If there was one thing that you could ask for – one improvement that you think would allow you to make a breakthrough in this research, what would that be?
To recreate the BBB, which shows such a tight barrier function to a variety of macromolecules and drugs of different sizes. That’s a very big challenge that would have a very significant impact in improving the tumor model. Another thing to be improved is that our model is still very simple. We do not have the immune components – essentially microglia – that have been shown to play a very big role in glioblastoma remission and immune response. It will be very significant if we could integrate these components into the model.