Human herpesvirus 6: the ancient herpes
Human herpesvirus 6 is a common childhood infection, but for some people, the virus is inherited through the germline from a 24,000-year-old ancestor.
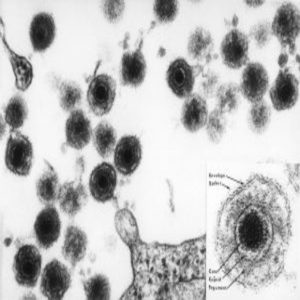
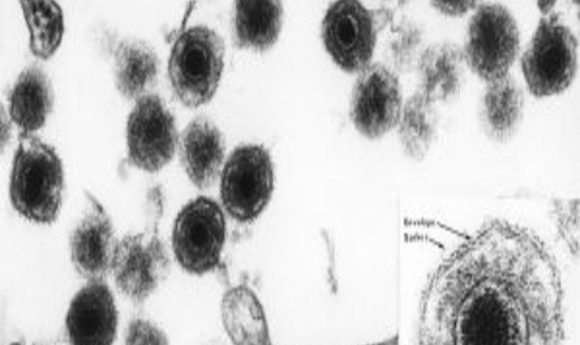
An electron micrograph of human herpesvirus 6 (HHV-6), including an insert of the mature virus particle. Like most herpes viruses, it has an “owl eye appearance.”
Credit: NIH Image Library
Before kindergarten, approximately 90% of children become infected with human herpesvirus 6 (HHV-6), more commonly known as sixth disease or roseola. Typically, it causes a 3-day fever, followed by a 2-day full body rash. Then like most herpes viruses, the symptoms clear and the virus enters a state of latency.
Human herpesvirus 6 (HHV-6) is commonly shared through saliva, but for an estimated 1% of people worldwide, the human herpesvirus 6 (HHV-6) genome is integrated into the telomeres of their chromosomes and thereby inherited through the germline. A new study in the Journal of Virology is the first to show that a seemingly unrelated population of Europeans with integrated human herpesvirus 6 (HHV-6) all inherited the virus from a common ancestor who lived 24,000 years ago.
“Until we published this study, there was really little understanding about what was going on,” said Nicola Royle from the University of Liecester, who is the senior author of the study. “What we didn’t know was how often the virus is getting to the germline and integrating. So, we decided to sequence viruses in individuals that had clearly inherited copies.”
Integrated human herpesvirus 6 (HHV-6) can be inherited as two different, but highly similar versions, HHV-6A and HHV-6B. Royle and her team screened 3,875 DNA samples donated to international research panels by people worldwide and identified 58 samples with chromosomally integrated HHV-6. They sequenced HHV-6 in 28 samples, 7 with integrated HHV-6A from England, Japan, and Scotland, and 21 with integrated HHV-6B from England, Italy, Scotland, Pakistan, and China.
They next compared the sequenced genomes of HHV-6A and HHV6-B to reference genomes from non-integrated HHV-6A and HHV-6B respectively. They found that the integrated genomes for each subtype were more similar to each other than to non-integrated versions of each subtype. For HHV-6B, the integrated HHV-6B genomes of Europeans were more similar to each other than to the chromosomally integrated HHV-6B genomes from China or Pakistan.
This led the researchers to investigate if these highly similar integrated HHV-6B genomes originated from a common ancestor. They isolated and sequenced fragments of DNA containing the junction between the human chromosome and the integrated virus genome. Using the base substitutions among the HHV-6B genomes that share the same junction fragment, they estimated that for a subgroup of Europeans from England, Orkney, and Italy, HHV-6B likely first integrated into the germline of a common ancestor approximately 24,538 +/- 10165 years ago.
According to Alex Greninger, an HHV-6 researcher at Fred Hutchinson Cancer Research Center who was not involved with the study, it is important to note that the ancient common ancestor identified in this study only pertains to this subpopulation, and that other populations may in fact have different chromosomally integrated HHV-6 common ancestors.
Royle said she was initially surprised by the finding. “I guess we were all talking in the field like [HHV-6 germline integration] was happening a lot de novo,” said Royle. But now her results clearly show that HHV-6 germline integration is ancient in some cases.
For Alex Greninger, the findings were not at all surprising. His team recently published a similar paper investigating the occurrence of chromosomally integrated HHV-6 in an American population. They also found that seemingly unrelated people share very similar chromosomally integrated HHV-6 genomes.
Interestingly, Royle and her team found that the chromosomally integrated HHV-6 genomes were completely intact, suggesting that viral reactivation is possible. However, she noted that the possibility of reactivation might be quite low. “You’d have to remove the epigenetic marks that silence the virus and there are so many steps that we don’t understand in that,” she said.
“HHV-6 is a fun area to work in because nobody knows anything,” exclaimed Greninger. “It’s very open space and that’s what’s fun about it.”
Royle now plans to continue working on human herpesvirus 6 (HHV-6), focusing on the virus’s integration into the telomere. “I have to admit, I find this field really very fascinating,” she said.