Male brains vs. female brains: is the difference in the (cell) numbers?
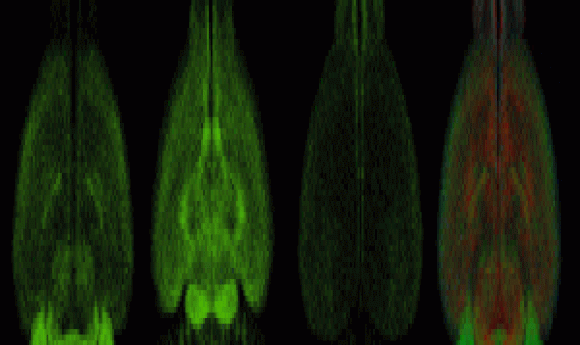
Scientists uncovered gender-specific differences in the brain, demonstrating the exciting power of the new qBrain brain-mapping platform.
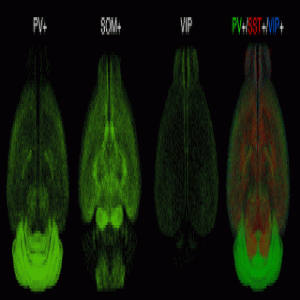
Fluorescent imaging of three primary inhibitory cell types in the brain.
Credit: Osten Lab, Cold Spring Harbor Laboratory
For years, scientists characterized isolated regions of the brain because technical limitations prevented analysis of the whole brain at once, requiring researchers to extrapolate findings from one region to another for similar functions. Pavel Osten and his team at Cold Spring Harbor Laboratory have now fixed this problem by designing a novel quantitative brain-wide mapping platform called qBrain. For their recent Cell paper, they employed qBrain in a series of proof-of-principle experiments and uncovered surprising results, including gender-specific differences in the numbers of neuronal cells in subcortical regions of mouse brains.
“It could have turned out to be a lot more boring, in a way,” reflected Osten, the senior scientist on the project. “We found things that we did not expect, and that gave us optimism that [qBrain] will do more comprehensive [analyses] across a much larger set of cell types; it will give a much more comprehensive understanding of the compositions of the individual brain regions with respect to the pieces that make the circuits.”
qBrain combines automated serial two-photon tomography to image the whole brain with automated computational algorithms to detect and count fluorescently labeled cells. Scientists then upload the generated data to an open online database. Each qBrain experimental data set includes cell counts, densities, and images of serial sections to visualize the distribution of cells along with other data.
To demonstrate qBrain’s capability, Osten’s team assessed the distribution of several primarily inhibitory neuronal cell populations, including cells positive for somatostatin (SST), parvalbumin (PV), and vasoactive intestinal peptide (VIP). To study these cells, the researchers fluorescently labelled them and let the qBrain pipeline do the rest of the work.
“Whatever is measured, that is what it is. There is a very low error, in fact,” explained Osten.
Despite being generally smaller than male brains, female mouse brains showed higher numbers of SST+ and VIP+ cells in brain regions corresponding to reproductive behavior. In male mice, only the posterodorsal preoptic nucleus, a region affiliated to the male-specific function of ejaculation, exhibited an increase in SST+ cells. Osten explained that the latter is almost a positive control for qBrain; the platform independently detected this region of the male brain—a region highly specific for male sexual function—that contains 3× more cells than the comparable area in the female brain.
“I really did not expect that we were going to find anything [in male–female comparisons],” explained Osten. “We did it because NIH now requires it.”
This whole-brain mapping approach also demonstrated that ratios of SST+, PV+, and VIP+ cells varied across cortical regions of the brain. Enrichment of certain cells correlated with a region’s function, suggesting that the brain evolved specific neuronal circuitry fitted for these specific functions.
“[The findings are] quite unexpected and I think really just a foundation of what we can do with this kind of technology,” said Andrea Beckel-Mitchener from the National Institute of Mental Health (ORDGMH), who did not participate in the study but served as program officer for Osten’s NIH-funded project.
Osten’s team now plans to image around 20 cell types a year for the next 5 years and to expand to other organisms beyond mice. They already have collaborations set up to study blind mole rats, prairie voles, and human brain tissues.