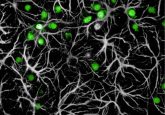
Simultaneously mapping genomes of thousands of single cells
Researchers have developed a quick, efficient technique for mapping the genomes of thousands of single cells. This high-throughput method may increase clinicians’ ability to measure genomic heterogeneity within tissues, paving the way for precision medicine applications.
Single-cell genome sequencing has proven valuable for the detection of somatic variation, particularly in the context of cancer. However, constructing sequence libraries for large populations of single cells using current technologies can be prohibitively expensive. The result is that the heterogeneity within tissues such as tumors is often overlooked.
A new low-cost technique from scientists at Oregon Health & Science University (OHSU) called single-cell combinatorial indexed sequencing (SCI-seq) provides a means of simultaneously generating thousands of low-pass, single-cell libraries for the detection of somatic copy-number variants.
“Traditional, low-throughput methods require you to sequence each cell very deeply,” said Andrew Adey, senior author of a new paper in Nature Methods describing SCI-seq. “But there’s an alternative strategy where you can do a lot more cells with lower coverage, allowing you to identify much more rare cell populations.”
Adey was among a group that recently established CPT-seq, a method to produce thousands of individually barcoded libraries of linked sequence reads using a transposase-based combinatorial indexing strategy. In combinatorial indexing, nuclei are barcoded multiple times before sequencing.
Adey figured that that same approach could be applied to single-cell genome sequencing. The key hurdle was removing nucleosomes bound to genomic DNA without compromising nuclear integrity. His team tested two approaches to removing nucleosomes: lithium-assisted nucleosome depletion (LAND) and cross-linking with SDS (xSDS).
Using a combination of LAND and xSDS, the researchers constructed sequence libraries for 16,698 single cells from a combination of cultured cell lines, primate frontal cortex tissue, and 2 human adenocarcinomas. They found that SCI-seq worked well for each cell type.
“Our technique will allow more precise treatment by targeting specific cell subpopulations,” said Adey, who is currently collaborating with clinicians at OHSU’s Cancer Center to deploy the SCI-seq technique.
“The other great thing about this technique is you can do it without any specialized, $100,000 pieces of equipment,” added Adey. “Pretty much all you need is access to a flow-sorting facility.”
In the future, Adey and his colleagues hope to use their technique to assess epigenetic properties of single cells, such as DNA methylation, that vary greatly between different cell types in the body.