Revealing the regulatory remnants of our Neanderthal heritage
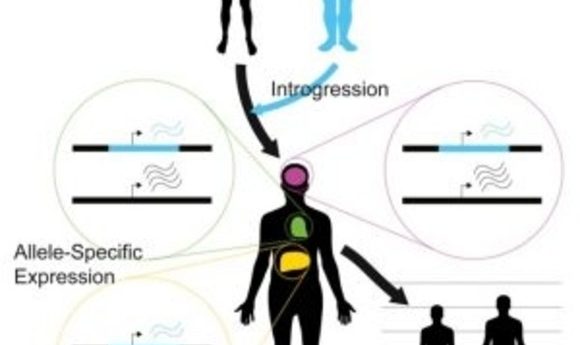
Neanderthal-inherited sequences are not silent remnants of ancient interbreeding, but have measurable impacts on gene expression that contribute to variation in modern human phenotypes.
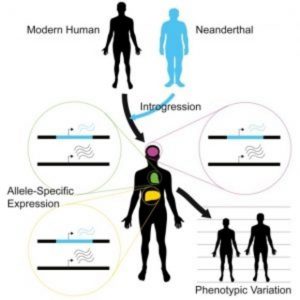
Before Neanderthals disappeared from the fossil record approximately 40,000 years ago, they interbred with ancestors of present-day humans. The legacy of this gene flow persists through Neanderthal-derived variants that survive in modern human DNA. The implications of this inheritance from the ancient hominin lineage most closely related to modern humans have not been entirely clear. To address this question, researchers recently developed a statistical approach and applied it to RNA-seq datasets to compare expression differences between Neanderthal and modern human alleles.
“We knew that Neanderthal sequences exist in the genomes of many modern human individuals, but before our study, it was unclear how biologically significant they were,” said Joshua Akey of the University of Washington. “Our study is important because it provides the first comprehensive look into how Neanderthal sequences influence gene expression variation in modern humans, which we showed were pervasive.”
Regulatory variation influencing gene expression is a key contributor to phenotypic diversity, both within and between species. While sophisticated methods have been developed to detect allele-specific expression in a single sample, across individuals or across tissues, researchers lacked a generalized approach to combine all information simultaneously.
To address this issue, Akey and his team developed a flexible Bayesian statistical approach to quantify allele-specific expression in complex RNA-seq datasets. As reported in the journal Cell, the researchers applied this method on a genome-wide scale to 214 individuals and 52 tissues, identifying widespread expression differences between Neanderthal and modern human alleles.
Perhaps most strikingly, Akey and his team found that the testes and brain regions including the cerebellum and basal ganglia showed significantly lower expression of Neanderthal alleles relative to other tissues. These brain regions have traditionally been associated with motor control and perception, but they also play a broader role in behavior and cognitive function—including language processing. Intriguingly, the cerebellum has undergone rapid expansion in the great ape lineage, and modern humans possess proportionally larger cerebella than did Neanderthals.
One brain-specific gene that exemplifies this pattern of downregulation is NTRK2, which encodes a protein that regulates neuron survival and differentiation as well as synapse formation. Mutations and polymorphisms in this gene have been associated with a range of neuropsychiatric and neurological disorders, including depression, suicide, impaired speech and language development, severe obesity, autism, obsessive-compulsive disorder, Alzheimer’s disease, anorexia nervosa, and nicotine dependence.
“As gene expression variation is known to have pervasive effects on phenotypic variation and disease susceptibility, our study suggests that DNA sequences inherited from Neanderthals also likely influence variability of many traits and diseases,” Akey said. “Importantly, previous studies, including our own work, suggested Neanderthal sequences were associated with various traits and diseases, but our study suggests the mechanistic basis of these associations is likely to involve gene regulation as opposed to changes in protein structure.”
According to the authors, the study demonstrates that Neanderthal-inherited sequences are not silent remnants of ancient interbreeding but have measurable impacts on gene expression that contribute to variation in modern human phenotypes. “Our study provides an important step in better understanding the functional, phenotypic, and evolutionary significance of sequences inherited from Neanderthals,” Akey said.