Does junk DNA hold the key to aging?
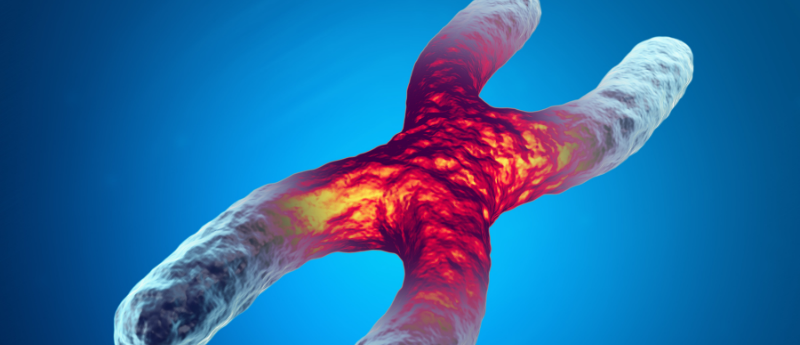
Researchers have discovered a DNA region that drives the activity of the telomerase gene, a gene that influences the aging of certain cell types.
Researchers from Washington State University College of Pharmacy and Pharmaceutical Sciences (WA, USA) have identified a DNA region, known as VNTR2-1, that drives the activity of the telomerase gene – the gene that encodes the telomerase enzyme, which is associated with the aging of certain cell types.
Telomerase is an enzyme that adds repeat sequences to the 3’ end of telomeres – regions at the end of each DNA strand, which act as protective caps to prevent chromosome damage.
In normal cells, telomeres shorten with successive rounds of DNA duplication and cell division, and eventually, the telomeres become so short that the cells can no longer reproduce, causing them to age and die. However, in certain cell types, such as reproductive and cancer cells, the activity of telomerase ensures that the length of telomeres is maintained. This restarts the aging clock in new offspring and is the reason that cancer cells are able to multiply and form tumors.
Determining how the telomerase gene is regulated and activated could be key to understanding how humans age, as well as developing novel cancer treatments.
The discovery that the VNTR2-1 region is involved in the regulation of the telomerase gene is particularly notable because of the type of DNA sequence it represents.
“Almost 50% of our genome consists of repetitive DNA that does not code for protein,” Jiyue Zhu, lead author, commented. “These DNA sequences tend to be considered as ‘junk DNA’ or dark matters in our genome, and they are difficult to study. Our study describes that one of those units actually has a function in that it enhances the activity of the telomerase gene.”
 Vindication for a misunderstood polymerase
Vindication for a misunderstood polymerase
In the latest shake-up for scientific dogma, research has demonstrated that eukaryotic cells can convert RNA sequences back into DNA, through a misunderstood, ‘error-prone’ polymerase.
The researchers discovered the link between the VNTR2-1 region and the telomerase gene by conducting a series of experiments that found that deleting the VNTR2-1 DNA sequence from cancer cells — both in a human cell line and in mice — resulted in telomeres shortening, cells aging and tumors stopping growing.
Zhu and his team then turned their attention to surveying the length of the VNTR2-1 region in human populations. DNA samples were collected from Caucasian and African American centenarians and control participants in the Georgia Centenarian Study, a study that followed a group of people aged 100 or above from 1988–2008. The results showed that the length of the sequence varied greatly throughout the population, ranging from as short as 53 repeats of DNA, to as long as 160 repeats.
“It varies a lot, and our study actually shows that the telomerase gene is more active in people with a longer sequence,” Zhu explained.
The team found that very short sequences were only identified in African American participants, and then on top of this, they found that there were relatively few centenarians with a short VNTR2-1 region compared to control participants. Zhu noted that the length of the sequence does not necessarily dictate lifespan, as a less active telomerase could make you less likely to develop cancer.
“Our findings are telling us that this VNTR2-1 sequence contributes to the genetic diversity of how we age and how we get cancer,” Zhu commented. “We know that oncogenes – or cancer genes –and tumor suppressor genes don’t account for all the reasons why we get cancer. Our research shows that the picture is a lot more complicated than a mutation of an oncogene and makes a strong case for expanding our research to look more closely at this so-called junk DNA.”
The study, published in PNAS, highlights the importance of junk DNA, and has the potential to deepen our understanding of aging and cancer.