A tale of two stem cells
A new strategy enables researchers to easily differentiate the dynamic states of pluripotency in stem cells.
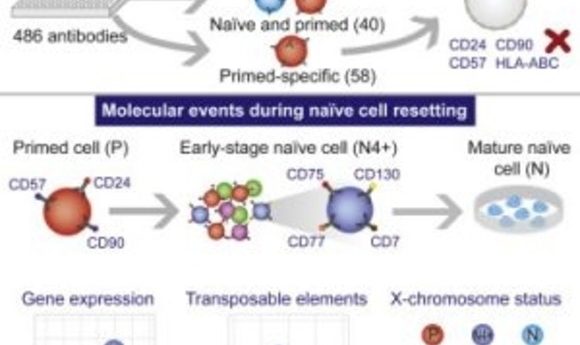
The ability of human pluripotent stem cells (PSCs) to give rise to all of the cells in our body, from neurons to kidney cells, has fascinated researchers in developmental biology and regenerative medicine. PSCs exist in two dynamic states, naïve or primed, and recapitulate an early or late stage of embryonic development, respectively. However, researchers cultivating PSCs in the lab are unable to easily identify whether a cell is in a naïve or primed pluripotent state due to substantial heterogeneity of the cells in culture, preventing any careful investigation into these specific cell types and their functions.
“We needed a way to pull out the right cell types at the right time,” said Peter Rugg-Gunn, senior author of a new paper published in Cell Stem Cell that describes a novel method for identifying markers that are specific to a particular pluripotent state.
To develop the method, the researchers cultured PSCs in different naïve and primed states and then exposed the cells to an exhaustive antibody screen targeting almost 400 cell-surface proteins. Next, they sorted and analyzed the cells by high-throughput flow cytometry, after which they identified distinct gene sets that are dynamically altered during naïve or primed states. “Once we had this list, there was then an awful lot of work involved to characterize and perform a substantial amount of validation using different lines and different culture methods,” said Rugg-Gunn.
In the lab, researchers can reset primed PSCs to a naïve state using a variety of culture techniques. However, this method is often variable and ineffective. Using these state-specific cell markers, the researchers captured this population of naïve cells at a stage in the resetting process that was much earlier than previously possible.
“I think this opened up a lot of exciting opportunities,” said Rugg-Gunn. The researchers went on to provide the first look at transcriptional dynamics during conversion from the naïve to primed states. “The transcriptional data allowed us to infer information about certain events including the dynamics of X chromosome reactivation during the reprogramming process.”
With the antibody panels and cell-sorting strategy now in place, Rugg-Gunn and colleagues would like to revisit their initial question: What epigenetic changes are occurring in cells that are transitioning between naïve and primed pluripotent states?