The Microsetta Initiative in the time of COVID-19
The extent of diversity associated with the human microbiome is vast, and largely uncharted. While there has been an impressive investment in the exploration of the microscopic in recent years, it is conservative to say we’ve barely made a dent in mapping the complexity of microbes associated with humans, let alone those of our furry compatriots.
In an effort to rapidly expand our understanding of the human microbiome, the Knight lab (University of California, San Diego [UCSD], USA) and collaborators created the American Gut Project (AGP) in November of 2012, on the heels of the massive $173 million NIH Human Microbiome Project (HMP). The HMP, while impressive in scale at the volume of data, was small in the number of humans who were able to take part at roughly 250 clinically healthy individuals in their 20s and 30s. This was a major limitation: the vast majority of humans on the planet were not eligible to participate, biasing our understanding of how microbes relate to health. What’s more, as these individuals were clinically healthy, the HMP data could not afford direct insight into the intricacies of how microbes influence disease. The AGP addressed this by providing a platform through which virtually anyone could participate, and contribute their stool to science. Of course, the AGP was initially still limiting as the participants generally had to pay to be part of the study, and it biased against those with an immediate interest in microbiome research in the USA. However, as interest in the project grew, partnerships in the UK and Australia were formed to make it easier for people in those countries to participate; additionally, groups began to sponsor studies to enable specific subsets of the population (based on disease type and socioeconomic status) to participate. Projects overseas, though, had to bank samples and cover the hideously expensive bulk shipment on dry ice.
In May of 2018, after the AGP published its first major paper examining samples from over 10,000 people, the main collaboration moved toward a grander vision: an inclusive open-access framework by which anyone in the world could participate called The Microsetta Initiative (TMI). While 10,000 is a HUGE amount of samples, the population was heavily weighted to just a few countries, with almost certain biases in those countries in part due to the cost of participation. It’s now well recognized that humans differ a lot by geographic region, and the reasons are not well understood. The major barriers for participation in the AGP were recognized as language, access and cost. TMI set out to lower these barriers, to gather a more comprehensive open-access database of microbial configurations, and to enable fundamental questions about what aspects of microbiomes are invariant across cultures. Central to the TMI is the question: how do results translate among the heterogeneous lifestyles, diets and health seen around the world?
Over the last 2 years, TMI has formed new partnerships to enable everyone to contribute to microbiome science. This has resulted in working toward the translation of documentation and the design of a language agnostic website as well as performance of a wide array of experiments to design a new collection kit that is non-toxic, complies with international shipping regulations, does not require cold chain shipping and is cheap, but still suitable for shotgun metagenomics and untargeted mass spectrometry. The aim was to rapidly expand our knowledge of microbes in multiple dimensions, and just like the AGP from which it grew, for the data to be released de-identified into the public domain for anyone to use.
 Talking Techniques: Donald Ingber on COVID-19, organ-on-a-chip technology and the Wyss Institute
Talking Techniques: Donald Ingber on COVID-19, organ-on-a-chip technology and the Wyss Institute
In the first installment of this two-part episode with Donald Ingber, Founding Director of the Wyss Institute (MA, USA), we discuss his invention of organ-on-a-chip technology, how he is utilizing them in the fight against COVID-19 and the Wyss Institute’s response to the pandemic.
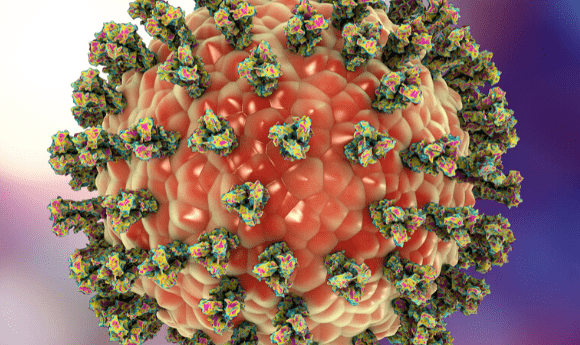
Then, just as TMI had geared up to produce tens of thousands of kits for distribution, SARS-CoV-2 struck
Like so many around the world, we were immediately determined to do our part to help reduce the spread and provide information about the epidemiology of the disease. Our team at UCSD first began partnering with the wider San Diego Coronavirus Response efforts — a collection of labs from multiple institutions collaborating on research, instruments, supplies, personnel and protocols. Our main focus was to develop a research-grade screening method for the detection of SARS-CoV-2 that critically does not compete for supplies with the CDC clinical test, but which has parity in results. As part of this effort, it was necessary to explore what types of buffers are suitable for the detection of the virus. The buffer used for the TMI kits, 95% ethanol, deactivates the virus making it safer to handle than live virus (although still requiring BSL2+ laboratories). In addition, while not specifically designed to fix RNA, the buffer has shown to be reasonable at retaining viral RNA, and sufficient to allow RT-qPCR detection as well as viral genome sequencing.
Detection of the virus is only part of the story though. Observing the virus does not provide information about antibodies developed. Through generous support from Becton Dickinson (NJ, USA), we obtained a large number of blood lancets for skin pricks. In parallel, and in coordination with the Human Research Protections Program at UCSD, we obtained approval to include these blood lancets in the collection kits, positioning the collection kits to provide comprehensive detection of the virus over multiple body sites (fecal/oral/skin); and the ability to examine seroconversion over a cohort, in a kit suitable for high throughput microbiome sequencing, creating the potential to ask whether variation in symptoms that COVID-19 positive individuals present is related to microbes.
We are now looking to collaborate with principal investigators who have existing cohorts and that are eager to gather microbiome and SARS-CoV-2 data. Participation is possible either under our IRB, or an existing one. We need your help to get our April production run, of over 20,000 kits, out to well the varied cohorts. The time to make an impact is now.
To learn more, please visit our website at https://microsetta.ucsd.edu or contact us at [email protected].
If you would like to keep up to date with our content on coronavirus, you can sign up for our site here, where you can subscribe to our newsletters for free!