Amoebae help discover new tuberculosis drugs
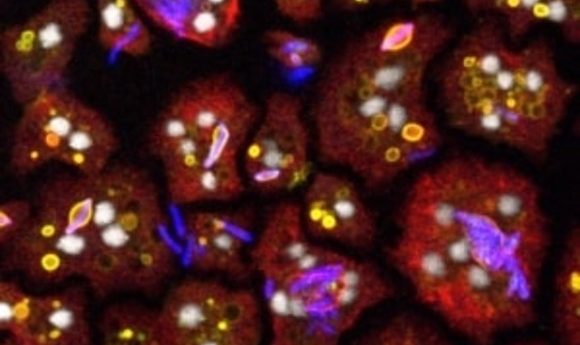
To screen anti-TB compounds, scientists used a simple host—free-living amoeba.
Drug design is a complex process, and it is particularly challenging to find drugs against manipulative and effective human pathogens. Many properties of tuberculosis (TB) bacteria, including a slow growth rate, impenetrable cell wall, and the ability to hide in granulomas inside the host lungs create roadblocks for the drug discovery process. To overcome these difficulties, scientists are now looking at alternative pathogen and host substitutes. In a new study published in Scientific Reports, researchers used a unique and previously under-appreciated host-pathogen model system—free-living amoebae and Mycobacterium marinum—to screen potential anti-TB compounds.
The drug company GlaxoSmithKline recently screened a library of 2 million compounds and reported 168 representative anti-TB candidates. Thierry Soldati, a microbiologist from the University of Geneva, Switzerland has been working on host-pathogen interactions for a long time and was interested in learning how the compounds from the GSK-TB set behave in living infection systems.
Amoebae are emerging as ideal choices for studying the course of various infections because they are easy to manipulate, generate few ethical concerns, and show promise in both early drug discovery and future target verification. Furthermore, M. marinum, a fish pathogen and a relative of Mycobacterium tuberculosis, is also emerging as a safer choice for conducting TB related research.
“Mycobacterium is an intriguing pathogen, and we wanted a simple system to screen anti-TB compounds—a system that is versatile and cheap, yet effective,” said Soldati.
Soldati’s team first explored anti-bacterial activities of the GSK-TB compounds against M. marinum in the absence of the host. They found that the majority of the compounds decreased the growth of M. marinum in culture. The authors then set out to determine if the drugs are cytotoxic for the host. For this, they used both amoeba and a mammalian macrophage cell line. They found that a majority of the drugs were not cytotoxic.
The authors next tested the anti-infective activity of the compounds against M. marinum by examining its growth inside amoebae and mammalian cells. They were surprised to learn that around 90% of the compounds didn’t show significant activity.
“Drugs were working on bacteria in broth but not in the host,” said Soldati. Overall, only 8 compounds showed strong anti-bacterial activity in the protozoan system and 19 in the mammalian infection system. Among moderately infective compounds, 21 showed similar effects in both hosts. In the two systems 13–14% of the compounds actually displayed pro-infective activity by increasing bacterial growth.
Soldati and his team further explored the mode of action of these compounds. For this, they performed mechanism of action studies to identify molecular targets of some of the anti-bacterial compounds. Based on those studies, the authors reported four new molecular targets.
Overall, the paper presents proof-of-concept for the M. marinum – amoeba system for anti-tuberculosis drug screening. The results show that although amoeba based assays are more stringent than the mammalian cell based assays, they can be used to predict the activity of compounds in infected mammalian cells.
“We are continuing to work on the proof-of-concept. The biggest obstacle now is to determine the targets,” said Soldati.