What are genetic algorithms?
Genetic algorithms borrow their name and principles from biological evolution, but can they help researchers discover the fundamentals of life?
Evolution is one of the most widely known theories in the world, and it’s not only because of the rich history of thought and ongoing debate about the origin of species. Some scientists believe the theory’s natural selection mechanics and inheritance principles are applicable to many fields outside of the life sciences.
One of those scientists, John Henry Holland, pioneered a class of search and optimization techniques called genetic algorithms (GAs) by borrowing terms and ideas from biological evolution. In his 1975 book, Adaptation in Natural and Artificial Systems, Holland explained how computational tools such as GAs play important roles in almost every field—especially those dealing with complex adaptive systems—from integrated circuits to image processing to economics to climatology (1).
The number of GA applications in the life sciences has exploded in recent years because of the amount of data researchers deal with on a day-to-day basis and the complexity of simulating certain biological systems. But do GAs truly follow principles of evolution and genetics to the T?
“I wouldn’t say that there is a ‘direct analogy’ between GAs and the natural evolutionary process, but they use an interpretation of the biological process to achieve their aim,” said Christos Dimopoulos, an associate professor of computer science and engineering at the European University Cyprus.
GAs fall under a larger discipline called evolutionary computation, which is a subfield of artificial intelligence. The main goal of these algorithms is to search for potential solutions to a given problem by following a simplified structure of evolution. The goal of GAs isn’t to look for a “perfect” solution to a problem, but rather to search for “fit” or sufficient solutions.
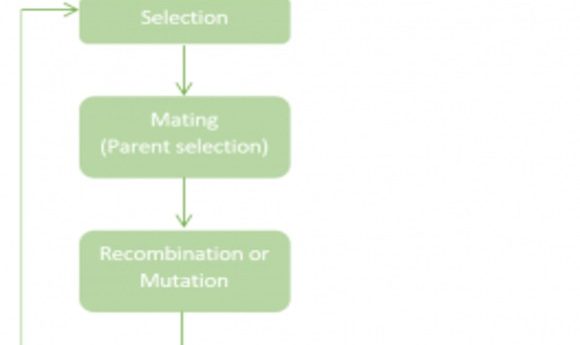
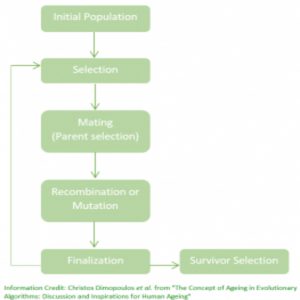
The process of a GA begins with an initial population of randomly generated solutions, where each individual solution is known as a “chromosome” and is represented by a sequence of 1s and 0s. Through selection, only the fittest individuals survive, which is determined by a “fitness function” that mathematically allows fit pairs to pass their solutions to the next generation. The selected parent chromosomes “mate” and produce “offspring.”
Genetic operators, such as “recombination” and “mutation,” occur at this point to introduce more diversity and produce better solutions in the next generation. These operators work similarly to biological principles; recombination preserves good parts of the genetic material or good solutions, and mutation introduces new genetic material or alternative solutions. After each generation, the GA does a final check to see if an optimal solution was found for the problem. If not, more generations will be produced until a sufficient solution is found.
“GAs are efficient and capable tools that can explore big search spaces and search for potential solutions, even when you don’t have a full understanding of the problem,” said Liucheng Guo, a Ph.D. student in the Electrical and Electronic Engineering department at the Imperial College London.
Guo believes GAs can be particularly useful in life science, and he’s currently working to develop better high-performing GAs. One such improvement is “migration” between parallel GAs—similar to gene flow—which allows for the exchange of solutions between two GAs. This technique can reduce execution time and increase the quality of chromosomes in each generation.
Many interdisciplinary groups around the world are beginning to use GAs for biological research by tailoring them for each specific project. A lot of those techniques adopt other principles of evolution as well.
Dimopoulos and his team recently published an analysis of biological aging compared to aging in GAs (2). In age-based selection for GAs, parents “die off” after a new generation of offspring solutions have been created, making the population of fit solutions smaller. This phenomenon is similar to the biological concept of “genetic drift,” where losing individuals in a population affects the genetic makeup of the group.
In terms of possible applications for age-related research, Dimopoulos suggests that GAs may be helpful in areas such as gene editing, stem cell research, and even cloning—at least as simulations to understand aging in terms of genetic damage. Some other important GA applications currently used in the life sciences include building phylogenetic trees, multi-sequence alignment, and protein predictions in protein interaction networks (3).
As the needs of scientists continue to grow when facing more challenging computational problems, bio-inspired algorithms may be an effective tool in understanding the fundamentals of nature.