Putting DNA fingerprinting behind bars

As potential developments in the automation of DNA fingerprinting threaten to introduce vulnerabilities in the system, researchers develop ‘genetic barcodes’ to prevent sample tampering.
DNA fingerprinting has become a cornerstone of forensic investigation since its initial development in 1984 and as a result of its popularity, the technique has experienced a flurry of advances to make it faster, cheaper and more widespread.
DNA fingerprinting relies on PCR to amplify the sequences from key sites in a DNA sample that are well documented to contain significant portions of the 0.1% of the genetic variation found between human genomes. This practice saves time and money by avoiding the need to sequence the entire genome, which currently costs over $1000, 99.9% of which will be identical between two individuals.
Many of the developments in DNA sequencing technology have been in developing smaller, automated and more practical devices to conduct the PCR stage of DNA sequencing, among other processes. However, with less human involvement the potential to corrupt or tamper with the process has grown, resulting in a new collection of vulnerabilities named “cyberbiosecurity threats.”
Searching for a way to fortify the technique against potential adversaries, a group of researchers from Duke University (NC, USA) and the New York University’s Tandon School of Engineering (NY, USA) have developed a method to insert a DNA barcode into DNA fingerprinting samples. This method will provide forensic scientists with confirmation that a sample is authentic and has not been tampered with.
 A silent witness: how analysis of pollen can solve crimes
A silent witness: how analysis of pollen can solve crimes
The specialized field of forensic palynology has been slow to garner worldwide momentum. With many advantages over other biological avenues of evidence, why is this technique so under-utilized and who are the giants of forensic palynology?
“Our idea is to inject non-harmful material into genetic samples immediately when they are collected in the field that act as a similar password. This would ensure that the samples are authentic when they reach the processing stage,” explained Mohamed Ibrahim (Intel Corporation, CA, USA), recently of Duke University.
The first step in the development of this protection was to produce two short sequences of synthetic DNA, 280 and 190 base pairs long. Each base of these synthetic sequences – or ‘barcodes’ – can be selected, meaning that each sequence contains a possibility of 4 different bases in each position, like a coded padlock with 280 or 190 wheels. These sequences can be added to the sample containing a potential perpetrator’s DNA.
To accompany these synthetic sequences, specific PCR primers were designed that would match the start and end sections of each sequence. These primers can be sent across to the forensic lab separately to the DNA sample. Used during the PCR step of DNA fingerprinting, the primers will lead to the multiplication of the barcodes, resulting in their display in the data resulting from the chosen identification method – for instance as specific bands in the Southern blot below.
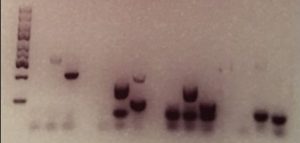
Credit: Duke University
As long as the barcodes appear as expected, then the sample can be confirmed as genuine. “When the right primers are used to unlock a barcode, you should get a positive result. If you don’t, then that means that the sample is not genuine. Some sort of switching or alteration has occurred,” explained Ibrahim.
A current vulnerability of this method of protection is the transport of the matching primers to the lab, but the team are already looking for ways to sure up the system. A potential solution could be to associate the barcode used with a certain aspect of the sample that will then indicate to technicians which primers to select from an available database.
No doubt, more vulnerabilities will be uncovered in the future, but with researchers continually working to protect these systems, hopefully innovations such as these genetic barcodes can help DNA fingerprinting to keep criminals behind bars.