Aptamers in disease research: moving beyond association and into action
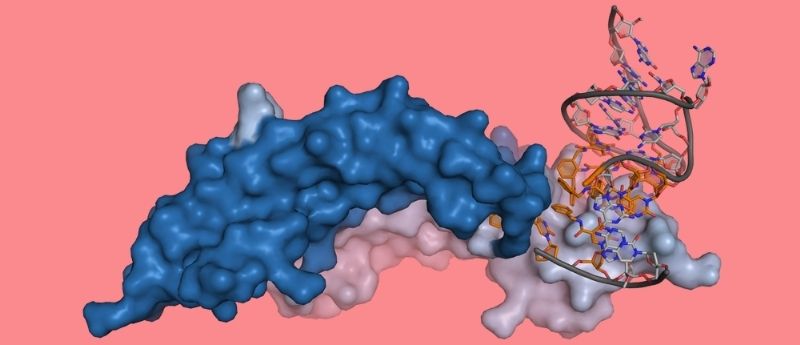
Recent developments in techniques for proteomic analysis, particularly in aptamer-based technologies, have led to a series of advances of several disease types, from infectious diseases to neurodegeneration.
The sophistication, speed and accessibility of genomic techniques have provided countless insights into disease risk and highlighted key indications for the biology behind specific diseases, with genome-wide association studies (GWAS) leading the charge. The resulting benefit to disease research cannot be understated; however, the full translational utility of these insights has not been realized. In many cases, we have not been able to convert the knowledge of a variant’s association with a disease phenotype into an understanding of the molecular mechanisms that produce it.
In part, this is due to the increased complexity and variability of the molecules downstream of nucleic acids that are integral to these mechanisms: proteins. Another contributing factor, however, is that the advancement of proteomic techniques has lagged behind next-generation sequencing in many respects. Well-established proteomic analysis methods, such as antibody assays and mass spectrometry, often exhibit throughput, dynamic range or operational times that can limit the scope of a study.
The advancement of more contemporary techniques has been in progress, particularly in aptamer-based screening and detection technologies, which depend on highly specific, synthetic single-stranded DNA molecules to bind to a target protein. Whilst aptamers have existed for over 30 years [1], recent advances in these technologies – improving the dynamic range and the number of proteins that can be screened – have enabled researchers to conduct more expansive studies of the proteins involved in numerous diseases states [2].
Decoding cardiovascular health
One such study recently examined two measurements typically associated with good health – baseline VO2max – and the capacity to improve this baseline through exercise regimens – ΔVO2max. An international collaboration, headed up by senior author Robert Gerszten of the Beth Israel Deaconess Medical Center (MA, USA) set out to establish the proteins that are associated with these two values and shed light on the pathways relating to cardiovascular fitness.
Using a SOMAscan assay, the team was able to measure the presence and abundance of almost 5000 proteins in the plasma of 650 participants. Participants were selected for their healthy, yet sedentary, nature and had their plasma assessed before and after a 20-week cardiovascular exercise program. Baseline VO2max was also measured at the start and end to establish ΔVO2max.
 Infographic: The future of proteomics
Infographic: The future of proteomics
Proteomic identification and detection have been carried out by mass spectrometry and antibody-based assays for a long time, but pioneering aptamer-based technologies now enable an unprecedented scale of detection.
The research found no correlation between each participant’s initial baseline VO2max and ΔVO2max, highlighting the limited ability of clinical traits, such as height and weight, to predict ΔVO2max. This finding was largely replicated in the team’s proteomic analysis; 147 and 120 proteins were associated with baseline VO2max and ΔVO2max, respectively, with only five proteins included in both groups.
Interestingly the team found that their results, whilst clearly highlighting why there is a minimal link between VO2max and ΔVO2max, could also suggest how these proteins could impact each characteristic. An association was established between a number of the proteins associated with VO2max and pathways related to hematopoiesis, angiogenesis and metabolic processes. ΔVO2max-associated proteins were noted to correlate with key signaling pathways, autophagy and extracellular matrix regulation.
Testing their findings, the team produced a protein score to predict the capacity of a participant to improve their VO2max. When combined with existing clinical traits known to have a minor predictive ability, this score allowed them to predict clinically significant improvements in VO2max with considerable success.
To assess the relevance of the identified proteins to overall health, the researchers investigated the proteins’ association with incident-all cause mortality. Thirty-six and 20 proteins associated with VO2max and ΔVO2max, respectively were also documented in the Framingham Heart Study; with twelve VO2max-associated proteins and nine ΔVO2max-associated proteins associated with incident-all cause mortality in the world-famous study. These findings highlight several proteins of interest for further studies to establish the mechanisms that link these markers of cardiorespiratory health to mortality and their value as plasma biomarkers for poor health.
 Modeling a heart attack: researchers develop human cardiac organoids
Modeling a heart attack: researchers develop human cardiac organoids
Researchers have developed human cardiac organoids in order to model myocardial infarction and drug cardiotoxicity, a significant development in the development of therapeutics for cardiovascular diseases.
Aptamers also play a more direct role in the examination of disease states. Patients with type 2 diabetes (T2D) have a significantly increased risk of coronary artery disease (CAD) and stroke, so much so that they account for almost 50% of the 15 million people who die of these conditions annually [3].
With this haunting statistic in mind, an Italian research collaboration set out to identify a more comprehensive list of proteins differentially associated with both T2D and CAD, with the aim of exposing new potential biomarkers and drug targets. The team, led by Ele Ferrannini (National Research Foundation Institute of Clinical Physiology, Pisa, Italy), applied aptamer-based technologies to analyze almost 5000 proteins in a cohort of 528 individuals. The cohort was separated into two groups. One group had diffuse lumen narrowing or plaques (CAD1), and one had clean coronaries (CAD2).
Of the top 20 proteins identified that showed a high association with either CAD or T2D, 15 were associated with both and several key proteins, including GDF15 and renin, were highlighted as potential biomarkers for the development of T2D and CAD and as therapeutic targets. What’s more, the well-established inverse relationship between adiponectin levels and T2D was shown to also be present with CAD.
In addition to indicating potential mechanisms behind the T2D and CAD relationship, the study was also able to show that for those with T2D, young females with lower renin levels were protected from CAD. This provides a clear clinical benefit for screening decisions.
Assaying Alzheimer’s with aptamers
Though protein-focused, aptamers can also play a useful role in mapping key aspects of the genome. By monitoring protein abundance with such accuracy, they can be used in combination with other methods to establish regions of the genome that influence protein expression.
In a study supervised by Oscar Harari and Carlos Cruchaga from Washington University (MO, USA), researchers used aptamers to establish the tissue-specific genetic controls of protein expression related to neurological conditions. To do this, they established a genomic atlas of protein quantitative trait loci (pQTL) – genomic loci that are responsible for changes in the extent of a protein’s expression – using a combination of GWAS and aptamer-based high-throughput screening technology.
Tissue samples of cerebrospinal fluid (CSF), brain and plasma from participants with and without Alzheimer’s disease (AD) were collected and the abundance of 713, 931 and 1079 proteins were analyzed in each tissue type respectively. This was coupled with a genome-wide association analysis to identify any SNPs or variants that were significantly correlated with changes in protein expression.
 A timeline of key Alzheimer’s disease milestones
A timeline of key Alzheimer’s disease milestones
What are the key Alzheimer’s disease milestones? Find out how far Alzheimer’s disease research has progressed using this interactive timeline. Read on to discover just how much there is yet to discover in Alzheimer’s research.
This approach identified 274, 127 and 32 pQTLs for CSF, plasma and brain tissues respectively and further analysis highlighted 23 proteins involved in AD risk across the three tissues. These findings were used to highlight potential therapeutics that could be repositioned for AD, nominate potential biomarkers for the disease and expose numerous avenues for further research into the basic understanding of the disease.
What’s more, the atlas, which can be found fully available online here, allows for detailed investigations into the pQTLs and their function. The authors were able to conduct a bioinformatic analysis to examine how these regions impact expression. Of particular note, the authors found that for cis-pQTLs there was a negative correlation between the distance to the transcription factor site and the strength of the association. This affirms the theory that most QTLs relate to transcription factor sites or chromatin accessibility and highlights the ability of the atlas to anchor signals obtained from GWAS onto functional genes.
It always comes back to COVID-19
Of course, no technique could prove its worth without pitting itself against the nemesis of our time, COVID-19. Just so, aptamers have played a part in detection and therapeutic development for the virus [4] and have revealed key details about the wildly variable response of humans on infection with the disease.
A recent collaboration set out to understand the genetic architecture responsible for the synthesis of proteins that interact with SARS-CoV-2 and control host reactions to the disease. The study was led by Aroon Hingorani (University College London, UK) and Claudia Langenberg (University of Cambridge, UK).
To do this, the team selected 409 human proteins that interact with SARS-CoV-2, are involved in viral entry or hyperimmune responses and are associated with disease severity. They then screened 179 of them that were available to examine using the most comprehensive aptamer-based proteomic assay, SOMAscan. This screen took place in samples taken from a cohort of 10,708 individuals prior to the COVID-19 outbreak.
 Sniffing out the source influencing COVID-19 severity
Sniffing out the source influencing COVID-19 severity
An exploration of the cells and expression patterns in the nasopharynx epithelium of COVID-19 patients has revealed a key component in the varying disease severity experienced by the infected.
Using a similar combination of GWAS and high-throughput aptamer-based assays employed by Harari and Cruchaga et al, the team identified 220 pQTLs for a total of 97 proteins. Of these, 45 proteins had never previously been associated with a pQTL, while 38 of them encoded existing drug targets. Further bioinformatic studies of the pQTLs allowed the researchers to investigate the potential impacts of targeting specific proteins
This study once again provides a readily available online tool for researchers to interrogate prospective drug targets and accelerates the identification of existing drugs that could be repositioned to fight the disease.
Due to their synthetic composition, aptamers are highly adaptable and replicable, traits that have enabled their applications to impact many more diseases. In cancer, aptamers have now been applied across the full spectrum of disease research, from diagnostics to therapeutics, where their high specificity and affinity make them attractive options for drug targeting [6].
In the field of disease research, which is crying out for a solution to help interpret the litany of genetic associations that still need to be explored, aptamer-based technologies are forcing their way into the fore as a vital tool for disease and drug discovery researchers.
This article was supported by Somalogic as part of our In Focus: Advanced proteomic analysis.