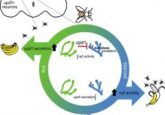
Could hibernation help to reduce obesity?

Researchers have used RNA sequencing to provide an insight into the underlying mechanisms of hibernation in grizzly bears.
Low activity in humans usually results in an accumulation of fat and consequential health deterioration, including an increased risk of Type II Diabetes and cardiovascular ailments. Bears, and other large hibernators, have evolved immunity to these effects during hibernation.
Consequentially, it is believed that by increasing our understanding of the physiological and cellular changes experienced in different hibernating species we could uncover a route of new treatments for human diseases, particularly those related to obesity and insulin resistance.
Researchers at Washington State University (WA, USA) have taken steps toward this by investigating the genetic changes in the Grizzly Bear (Ursus arctos horribilis) before and during hibernation.
“The number of differentially expressed genes is striking,” remarked Joanna Kelley, associate professor in the School of Biological Sciences and one of the co-authors on this paper.
In this study, published recently in Communications Biology, samples of liver, adipose and muscle tissue were taken from six bears during their active, hyperphagic (just before hibernation) and hibernation periods.
The researchers then extracted and sequenced RNA from these samples to determine any transcriptional changes between the different tissue types at the different points in their hibernation cycle.
- Sleep deprivation could increase risk of obesity
- The ancient canine cancer that has spread worldwide
- Have humans re-shaped dog brains?
During hibernation, the bears displayed dynamic gene expression changes in all three tissue types, but this change was greatest in the adipose tissue. This result was consistent with findings from studies using other mammals, though the number of changes was much greater in bears.
All three tissue types displayed similar changes during hibernation with an increased expression of genes associated with muscle protein anabolic pathways and reduced expression of genes associated with insulin signaling, urea production and muscle protein degradation.
During the active and hyperphagia periods there was a large variation in gene expression between the different tissue types. While there were many genes differentially expressed in the adipose tissue, there were only three genes in the liver tissue and none in the muscle tissue.
The researchers believe that the subset of genes discovered to be differentially expressed across all three tissue types could reflect a common regulatory mechanism for hibernation, though further research will need to be taken to confirm this.
Regardless, the gene families identified in this study could provide a useful option for developing novel therapies to treat human and animal diseases such as obesity, type 2 diabetes and atherosclerosis.