Where are the SARS-CoV-2 genomes from East Africa?

The need for SARS-CoV-2 genomes in managing the COVID-19 pandemic in East Africa, and the path to get them.
The first reported case of COVID-19 was 13 March 2020 in Kenya and 10 weeks later, not a single genome is available publicly from any of the East African Community countries (Kenya, Tanzania, Burundi, Uganda, Rwanda, South Sudan). Why is it so? And why does it matter? Globally the main focus during this outbreak has been rapid COVID testing and not whole-genome sequencing. The team at Nextstrain has highlighted the utility of whole-genome sequencing in addition to rapid testing. We have presented below some of the challenges to obtaining whole genomes in East Africa and most importantly we have suggested a way forward.
If you would like to keep up to date with our content on coronavirus, you can sign up for our site here, where you can subscribe to our newsletters for free!
Why are genomes important?
As a diagnostic, whole genomes are critical. Sequences confirm the identity of the disease-causing pathogen and can be further used for studying diversity, tracing movement of virus strains, designing models that can predict the disease spread and to better understand the enemy. A recent French study in bioRxiv has claimed the SARS-CoV-2 strain in France was not imported from China. This highlights the importance of a sequencing initiative to be able to properly trace the progress of the pandemic in every setting – the Icelandic approach.
Why is obtaining this data real-time as the pathogen is emerging important?
Real-time data are very important because they serve as a diagnostic test that guides quick patient management and decision-making from an epidemiological standpoint; and genomics would provide further tools in designing therapeutic approaches.
Sequencing facilities in East Africa
Over the years, millions of USD have been spent building genomic sequencing facilities in East Africa. In Kenya, Biosciences for east and central Africa (plant and animal) and KEMRI–Wellcome Trust (both Nairobi, Kenya) (human health) are partnerships with national governments and international funders but to date neither have delivered a genome.
In Uganda, the Uganda Virus Research Institute (UVRI; Entebbe, Uganda), is a centre of excellence in virus research with the human and infrastructural capacity and international support for genome sequencing. However, UVRI has also not yet delivered a single SARS-CoV-2 genome.
Tanzania has a different landscape. There are no large international sequencing facilities, but the national research organizations, universities and hospitals like Muhimbili National Hospital (Dar es Salaam, Tanzania) and the Sokoine University of Agriculture (SUA; Morogoro, Tanzania) have various platforms such as the Illumina (CA, USA) MiSeq, HiSeq and the Oxford Nanopore MinION. They too have not yet generated any SARS-CoV-2 genomes.
So why have none of these institutions with the sequencing infrastructure and support in Kenya, Tanzania and Uganda not delivered the much-needed SARS-CoV-2 genomes yet?
What are the possible causes of lack of SARS-CoV-2 genomes from East Africa?
- Is the sequencing capacity lacking- human and machines?
- Are the supplies for sequencing available in-country?
- Are computers available to do the bioinformatic analyses?
- Is money readily available to buy supplies?
- Is internet connectivity and bandwidth adequate to support bioinformatics and sequence analysis?
- Is there stable power to run the molecular labs?
- Are partnerships lacking in sequencing and diagnostics to carry out real-time tracking of new infections and spread?
- Is there lack of government support for whole-genome sequencing?
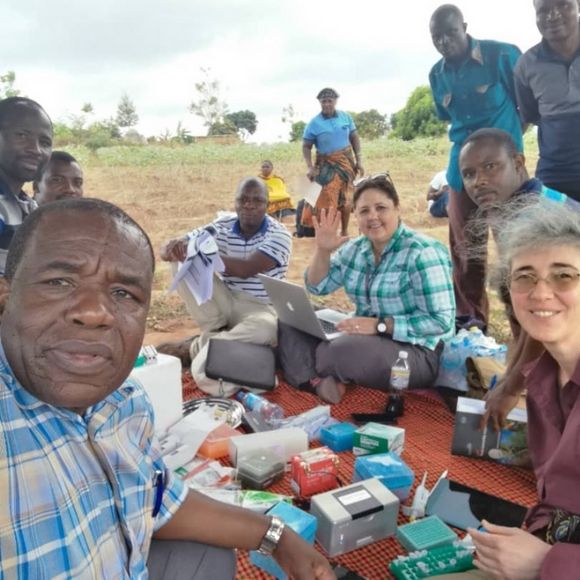 Taking the highest tech genomics tools to the farmers in East Africa
Taking the highest tech genomics tools to the farmers in East Africa
DNA sequencing in Africa is currently a laborious task requiring researchers to send data to a centralized sequencing lab in Kenya or to await results from overseas. Here, Laura Boykin tells her story of working with the Tanzanian Agricultural Research Institute.
Kenya
Lack of partnerships:
For Kenya, the biggest hurdle is a lack of partnerships. So far, all the work on COVID-19 is handled solely by the Ministry of Health (MoH; Nairobi, Kenya). Accordingly, there has been no access to samples – considering also that this disease is highly infectious and these samples need to be handled in biosafety level 4 labs. Due to poor partnerships (aka poor coordination), the work is largely being done in KEMRI and private medical labs such as Lancet. The other limitations are:
- Lack of funds for supplies to conduct the laboratory tests and sequencing, considering that this is a new disease.
- No sequencing capacity supported by the MoH; but this could easily be solved by outsourcing the services from other public institutions or the private sector.
- Limited government support – which is very closely related to 1 above. Government support and focus has only been to those involved in the health sector (hospitals and quarantine centers). I wish to add that it may not have been the government’s priority to spend its budget on sequencing,g perhaps because its importance has not been highlighted.
Power, computers, internet and PCR machines are not a challenge.
Tanzania
Sequencing capacity:
The sequencing capacity is there especially in research and academic institutes; the SUA has the Thermo Fisher Scientific (MA, USA) Ion Torrent that they use for foot and mouth disease and other animal research, the Kilimanjaro Clinical research Institute (KCRI, Moshi, Tanzania) has the Illumina MiSEQ (I have seen this personally) which they use for their tuberculosis research; the Government Chemist Laboratory Authority (Dar es Salaam, Tanzania) has a genetic analyzer and was able to acquire the Illumina HiSEQ, which they use for their forensic studies; the National Health Laboratory (Dar es Salaam, Tanzania) also has a genetic analyzer. There are two laboratories which are capable of sequencing using Oxford Nanopore Technologies (Oxford, UK). These are Muhimbili national hospital and the SUA in collaboration with the NHL. There were no funds to do the sequencing at the beginning of the outbreak but now the SUA has secured some funds to sequence, Muhimbili might get a donation to do so too. Another laboratory that is capable of sequencing but does not have the funds to do so is the KCRI. Capacity and skills are not a problem. However, in a government setting and in most institutes, employees are given specific tasks as per institute mandate. It’s true that we have many people trained in sequencing, but some are outside government settings/employment and some of those who are in government employment are not in clinical research. For example, the cassava disease diagnostic team was focused on agricultural research. Some of the trained people are not trained to handle clinical samples. So clearly there is a disconnect between clinical and agricultural disease diagnostic techniques.
Lack of partnerships:
Another challenge is lack of local partnerships (internal collaborations among different institutes in Tanzania). There are no good networks that connect healthcare facilities with research and academic institutes. Most healthcare facilities do not have the critical mass of trained experts in sequencing and due to their mandates and the sheer heaviness of their routine workload, they rarely have the bandwidth to pursue research regularly. Herein comes the need to forge strong links between the two that would have been in prime position to address this pandemic. Unfortunately it has not been easy; from my personal experience there are a lot of territorial issues at play that are hard to overcome. Perhaps this pandemic might bring a change in mindset.
Lack of funding:
Another challenge is global but is felt more in countries like Tanzania; inadequate funding for R&D. While the government, through the Tanzania Commission for Science and Technology (COSTECH) and other institutes, provides for R&D funding, it is still limited especially when compared to the costs of running genomics experiments. External funding is always difficult especially for researchers who are not part of a consortium led by PIs from Europe and/or North America. This has helped establish centers but has meant that the moment funding runs out the lab is less active, the reagents and consumables run out and equipment ends up in disuse.
Lack of awareness:
There appears to be a lack of awareness among policy makers and/or not enough initiative from the local scientists working in this field to inform our policy makers about the importance of whole genome sequencing for management of COVID-19. Since most sequencing initiatives in the country are led by foreign consortia (which we feel needs to change) led from either Europe or North America it is possible that the benefits from such projects are rarely seen by policy makers in Tanzania. We see there needs to be a clear link between the governments and local scientists to work on the same matters from different perspectives. We hope the donated research reagents to the African CDC will reach the institutes as soon as they arrive the airport without customs delays.
Uganda
Sequencing capacity:
There is both human and infrastructural capacity in sequencing at UVRI and the Medical Research Council all based at Entebbe, Uganda. However, the COVID-19 genomes are not yet out in the public arena.
Computers and supplies:
There are computers, access to internet, power and the supplies required to carry out PCR testing and analysis of coronavirus/COVID-19 infections, which were initially provided by the UVRI through its running projects and currently with the support of the government. However, more supplies would be needed to monitor the entry and spread of the virus in the communities.
Lack of partnerships:
As of today, it is the sole responsibility of the Ministry of Health (Kampala, Uganda) as the mandated institution of government to lead all COVID-19 pandemic-related issues. This includes checking for possible cases with suspected symptoms, isolation/quarantine, collecting samples, sample analysis and announcement of outcomes of testing and treatment. In addition, task forces were established to coordinate COVID-19-related issues at national, regional and district level. The laboratory analysis of the suspected COVID-19 samples is carried out by UVRI. Although there are other institutions with both human and infrastructural capacity in molecular biology and disease diagnostics, there are limited partnerships on widening the testing for COVID-19 in the country to involve the private sector. This may be partly due to the highly infectious nature of the disease and the requirement to carry out the laboratory testing and analysis in a biosafety level 4 containment facility such as UVRI. However, there are some partnerships within the private sector in management of the disease.
 Insight into SARS-CoV-2 genome spells good news for vaccine development
Insight into SARS-CoV-2 genome spells good news for vaccine development
Infectious disease researchers have identified just five SARS-CoV-2 gene variants, suggesting a vaccine for COVID-19 could be highly effective.
Suggestions for the way forward
- Organize funds for national labs to readily respond to new COVID-19 outbreaks and to monitor the spread of the virus in each country.
- Invest in portable diagnostics and genomics that can be deployed to the borders and regional centers.
- Stock supplies to do real-time PCR-based diagnostics and whole-genome sequencing in each country to ensure the timely detection of new disease pandemics such as COVID-19. The lack of industrial capacity for reagents and consumables in East Africa means the current lockdown on international air travel has magnified the already significant logistical challenges since most of these have to be imported. Road transport is slow and often too risky, where keeping temperature cannot be maintained for the heat sensitive reagents.
- Establish rapid testing and portable diagnostic units as management strategy in government responses to the emerging pandemics.
- Build and/or strengthen local and regional (East African) partnerships and collaborations to bring together all the expertise in molecular diagnostics and genomics in the public and private sectors in each country. We must do science together and STOP the ‘5-star hotel post-it note meetings”.
- Initiate a review of large internationally funded genomic research centers. Would it be better to empower local labs and not big regional hubs? These centers have not produced real-time whole-genome data that would have been critical in management decisions.
- International partners, such as universities who partner with national laboratories must be more inclusive in development of research capacities to include the human health and agricultural (crop, animal, fisheries and forest) sectors.
- Sequencing companies, such as Oxford Nanopore and Illumina, should invest in hiring African scientists. Not only can they build local communities, but the local knowledge will help find ways to get the supplies to scientists in the country. Included in this is a need for inclusive writing on the packages and understanding that each country has different customs regulations. Some requiring pre-authorization before sending materials.
- East African Community governments should increase funding for this type of work. They must also prioritize facilitating critical research and this must include 1) competitive salaries for scientists; 2) laboratories with constant power and internet; 3) computers and data storage; and 4) mobile data for scientists to work from home should circumstance force labs to retain only the technical personnel. These governments must also cut the red tape at the airports and borders to receive molecular lab supplies. Customs regulations must be updated to include a strategy for pandemics.
- Ensure accurate, objective and responsible dissemination of knowledge and information regarding the disease and its management to the general public.
- Use gender inclusive approaches and prioritize the engagement of women at every level.
This article is written by East African Scientists and international partners who have been working for years on collaborative research projects, including The Cassava Virus Action Project, around managing emerging plant virus disease pandemics using novel molecular diagnostics and genomics. The team was disheartened to watch COVID-19 arrive and spread in East African countries, where they have successfully partnered to build capacity in rapid plant virus diagnostics and genome sequencing using novel portable technologies such as the Oxford Nanopore MinION, which have not been put to good use in the fight against the pandemic.
Professor Elijah Ateka – Molecular Biologist
Dr. Joseph Ndunguru – Molecular Plant Pathologist
Dr. Daniel Maeda – Molecular and Cellular Biologist (Health Focus)
Mr. Charles Kayuki – Molecular Biologist
Dr. Peter Sseruwagi – Molecular disease epidemiologist
Dr. Laura M. Boykin – Computational Biologist