Three rules for making a broad-spectrum antibiotic
Scientists have discovered key characteristics that allow molecules to enter and accumulate inside almost-impenetrable gram-negative bacteria. Will this translate to an arsenal of new drugs for tackling antibiotic resistance in a brand-new way?
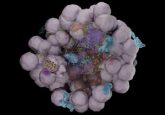
Surrounded by a lipopolysaccharide-studded outer cell membrane, gram-negative bacteria are tough to kill. Drugs can’t easily get into them, and the bacteria rapidly become resistant to the few that can. No new classes of gram-negative-active antibiotics have been approved since 1968.
Figuring out how to slip compounds into these cells, then, is key to winning the war against bacteria. A new study led by Paul Hergenrother at the University of Illinois reveals a novel set of rules for compounds that can enter and accumulate inside gram-negative bacteria; an antibiotic created using these rules slides through the bacteria’s porin channels in such quantities that efflux pumps can’t eject it before the cell is dead.
“What we wanted to do was go to a very basic level: instead of just doing what has been done unsuccessfully over and over, which is just screen for compounds that can kill some of these gram-negative bacteria, we wanted to try to figure out why such screens have been unsuccessful and then start to be able to use that information to create compounds that will kill these bacteria,” said Hergenrother.
Hergenrother’s group spent the past 5 years creating a cache of complex small molecules. They used a method called complexity-to-diversity, which takes complex natural compounds and chemically alters them one at a time in two to five steps. Their process produced nearly 200 difficult-to-purify compounds, containing elements such as primary amines and overlapping contiguous ring systems, that almost never show up in the huge screening libraries.
The team individually tested their compounds for the ability to accumulate in E. coli, a gram-negative bacteria, and found several that did. When they analyzed the data using sophisticated computational analyses, patterns emerged: molecules with primary amines, high rigidity (low rotatable bonds), and low globularity (less round, more flat) were more likely to accumulate. It made sense: molecules that are rigid, less globular, and polar could fit through porin channels more easily. These rules were different than the ones researchers had derived from retrospective studies examining similarities between gram-negative antibiotics.
Hergenrother and his colleagues then took an antibiotic against gram-positive bacteria, deoxynybomycin, that had two of the three characteristics of an accumulating molecule (low rotatable bonds and low globularity) and added the third, a primary amine, to it. They tested the new compound on E. coli, and not only did it enter the cells, it also killed the bacteria. Effectively, the researchers had converted a gram-positive antibiotic into a broad-spectrum antibiotic. The compound also kills several other types of problematic multi-drug resistant gram-negative bacteria.
Luckily, several gram-positive antibiotics exist that have two of the three characteristics that should allow them to enter gram negative cells.
“I’m hopeful that the logic and these predictive guidelines will be useful for a host of different compounds,” said Hergenrother. “We’re moving it forward in different ways, a lot needs to be done pre-clinically and just in general before one goes to a clinical trial. But we are marching down the road.”