An RNA binding agreement

Researchers have overcome the disadvantages of two different methods for identifying the target sequences of RNA-binding proteins by combining them into a new method.

Understanding the function of an RNA-binding protein (RBP) requires identifying its target RNA sequences. One approach for doing this is systematic evolution of ligands by exponential enrichment (SELEX), but this in vitro technique is not quantitative and favors the strongest binding sequences. Another approach is cross-linking immunoprecipitation (CLIP), but this in vivo method requires large amounts of cells and may select for highly abundant, low-affinity non-target sequences while missing low-abundance, high-affinity target sequences. In the July issue of BioTechniques, researchers led by Kee Kim at Chungnam National University describe how they overcame the limitations of these two approaches by combining them.
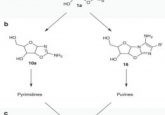
As with SELEX, their method uses RNA synthesized from a random oligonucleotide library, but the RNA contains the photo¬activatable ribonucleoside analog 4-thio-uridine that is used in CLIP. RNAs from the random sequence pool that bind the RBP are then UV crosslinked to the RBP and selected for by immunoprecipi¬tation of the RBP. This isolates low- and high-affinity sequences without being affected by target RNA abundance in the library pool.